New view reveals how DNA fits into cell
Cells are tidy packers, cramming DNA into nuclei to create a tangle-free, dense ball with pieces that are still accessible, researchers report October 9 in Science. The findings, based on a new three-dimensional view of the whole human genome, solve a long-standing biological mystery and may lead to a deeper understanding of how genes operate.
Except during division, a human cell's two meters of DNA is jammed into an area about a hundredth of a millimeter wide. But researchers had been puzzled by how cells could pack the DNA, which is organized into 23 pairs of chromosomes inside the nucleus, so tightly without hopelessly tangling it and making it impossible to use.
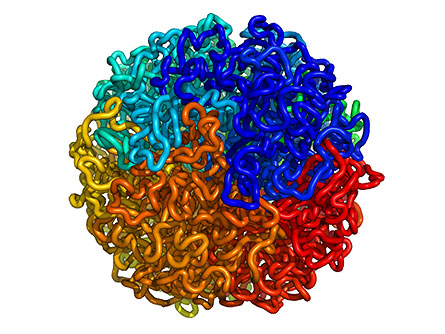
One of the reasons that DNA packing remained such a mystery is that scientists lacked the tools required to assess the shape of the entire genome. Earlier studies focused on the shape of small pieces of DNA that had been chopped out, removing them from their larger context. In the new study, Erez Lieberman-Aiden of Harvard University and MIT, Nynke L. van Berkum of University of Massachusetts Medical School in Worcester and colleagues developed a trick to lock pieces of neighboring DNA to each other while they were still in the nucleus. After removing the pieces and sequencing them, the researchers could calculate how close each and every piece of DNA had been to the other pieces and could reconstruct the 3-D shape of the genome.
"Our technology allows us to ask really fundamental questions about chromosomes," says study coauthor and molecular biologist Job Dekker of the University of Massachusetts Medical School in Worcester. "It really is a radical improvement over the previous technology. It's truly genome wide and unbiased."
Applying the method to human cells, the researchers found that the genome has a highly organized structure. Small pieces of DNA fold into globs, and those globs fold into larger globs and so on. The researchers report that this "globule of globules of globules" is fractal, meaning it is organized in such a way that it has the same pattern no matter how far you zoom in. This fractal shape is "super-dense, but has no knots," says Lieberman-Aiden.
Earlier studies by Alexander Grosberg, a theoretical physicist at New York University, first predicted the fractal structure of packed DNA. "Now this paper delivers beautiful confirmation of that prediction," he says.
The new analysis also found that the genome separates into two clear compartments: One is made up of stretches of DNA known to be active and working, and the other is made up of inactive DNA, set aside for storage, Lieberman-Aiden says. "The chromosomes are kind of weaving back and forth between those compartments," he says.
Future work with more comprehensive sequencing data may even allow researchers to discern individual genes. Part of the reason scientists are so intent on understanding the shape of packed DNA is that genes can be turned on and off by far-flung DNA elements, brought together by folding. By knowing which pieces of DNA are close to each other in the pack, researchers may be able to understand more thoroughly how genes are regulated. For example, misfolding on the large scale may disrupt proper gene regulation, which could lead to cancer, Dekker says.
"Now that we know the structure, we can ask questions like, why does it look like this?" Dekker also wants to understand how a gene and a regulatory element find each other in such a dense glob. As of now, "We simply don't know," he says.
Scientists also don't yet know whether this folding pattern holds true across different cells. Dekker says that there may be "a tremendous amount of variation."
Source: http://www.sciencene...dna_globule.jpg
Edited by Elus Efelier, 23 November 2009 - 04:46 AM.












































