.
F U L L T E X T S O U R C E : AgING
Abstract
To gain understanding on the mechanisms that drive immunosenescence in humans, we examined CD4+ T cells obtained from younger (20-39 years-old) and older (70+ years-old) healthy participants of the Baltimore Longitudinal Study on Aging (BLSA). We found that mitochondrial proteins involved in the electron transport chain were overrepresented in cells from older participants, with prevalent dysregulation of oxidative phosphorylation and energy metabolism molecular pathways. Surprisingly, gene transcripts coding for mitochondrial proteins pertaining to oxidative phosphorylation and electron transport chain pathways were underrepresented in older individuals. Paralleling the observed decrease in gene expression, mitochondrial respiration was impaired in CD4+ T cells from older subjects. Though mitochondrial number in both naïve and memory cells visualized with electron microcopy was similar in older versus younger participants, there were a significantly higher number of autophagosomes, many of them containing undegraded mitochondria, in older individuals. The presence of mitochondria inside the accumulated autophagic compartments in CD4+ T cells from older individuals was confirmed by immunofluorescence. These findings suggest that older age is associated with persistence of dysfunctional mitochondria in CD4+ T lymphocytes caused by defective mitochondrial turnover by autophagy, which may trigger chronic inflammation and contribute to the impairment of immune defense in older persons.
Introduction
Human aging is characterized by increased susceptibility to diseases and impaired capacity to handle stress and challenges from the environment, leading to excess morbidity and mortality [1, 2]. A critical factor in this process is a progressive decline of immunity, which results in reduced protection against external microorganismal threats, diminished surveillance and reduced repair capacity of damaged cells and tissues that can lead to degenerative diseases. Indeed, rates of hospitalizations with a discharge diagnosis of infection rise dramatically with age, from 74 to 86 per 10,000 in individuals age 20–39 years-old to between 432 and 891 per 10,000 persons for those who are 70 years-old and older [3].
Changes in cellular morphology and function that occur in the immune system with aging are globally referred to as “immunosenescence” and have been widely studied and discussed in the literature with evidence that they involve both innate and adaptive immunity [4, 5]. However, the core mechanisms leading to these changes are unknown [6]. Two contrasting phenotypes characterize immunosenescence, a status of chronically activated mild inflammation, which has been termed “inflammaging”, and a progressive reduction of the ability to mount an adequate immune response to vaccination and to infection [6]. Inflammaging is characterized by a robust increase with aging of blood levels of proinflammatory markers, and is a strong risk factor for the occurrence, progression, and complication of many chronic diseases, including cardiovascular and neurodegenerative diseases [7, 8]. Several aspects of the immune response are affected by aging, including proper initiation and, especially, maintenance and cessation [6].
Recently, it has been proposed that most, if not all the aspects of the aging process, may be attributable to a dysfunction of a limited number of molecular and cellular mechanisms with aging, that have been called the “Hallmarks” or “Pillars” of aging [9, 10].
The “Pillars” of aging paradigm has not been systematically applied to the study of immunosenescence. One aspect that is increasingly being recognized as important to explain the effect of aging on immune cells, particularly CD8+ T cells, is the role of one of the components responsible for cellular proteostasis, macroautophagy (hereafter referred to as autophagy) and one of its modalities, mitophagy [11]. Studies in CD8+ T cells of older individuals have shown lower basal autophagy levels [12], while autolysosomes accumulate in human CD8+ T cells undergoing replicative senescence “in vitro” [13]. In CD4+ T cells, it has also been reported that while autophagy activity is lower in cells from older individuals, it is preserved in cells from the progeny of centenarians, which correlates with improved function [14]. Experiments of T-cell-specific depletion of Vps34 [15] or Atg7 [16], essential genes for canonical autophagy, lead to accumulation of damaged mitochondria and ROS, showing that T cells depend on autophagy and mitophagy for homeostasis and function [17]. Several lines of evidence suggest that defective autophagy may impair the recycling of dysfunctional mitochondria and profoundly affects the functionality of T lymphocytes [18, 19]. Naïve T cells that encounter a new antigen undergo extensive proliferation and differentiate into effector T cells. The increased energetic demand is supported by a metabolic shift to aerobic glycolysis, also called the “Warburg effect” [20]. Mitochondria also contribute to this process by using TCA cycle metabolites for the building of macromolecules, including proteins and lipids as building bricks for new cells, and by producing ROS signaling required for full T cell activation [21–23]. At infection resolution, most T cells undergo apoptosis while a few remodel into memory T cells. In spite of low energy consumption at rest, memory T cells display a characteristic increase in mitochondrial mass, which creates a large reserve respiratory capacity that allows for rapid and sustained proliferation upon secondary exposure to antigen reactivation [24]. Under the hypothesis that age impairs mitochondrial function in lymphocytes because of defective autophagy, the resulting energy deficit may impair both the activation of naïve T lymphocytes and reactivation of memory lymphocytes upon a second encounter with the same antigen, which are both critical response mechanisms of adaptive immunity [25]. This is consistent with the blunted response to influenza after vaccination in older persons, and the finding that treatment with an analog of rapamycin (RAD001), which stimulates autophagy, effectively restores, at least in part, such a response [26, 27].
To gain further understanding on the causes of immunosenescence, we compared CD4+ T cells from younger donors (ages 20-39 years-old) and older (ages 70 years and older) participants of the Baltimore Longitudinal Study of Aging (BLSA). In quantitative discovery proteomics of cytoplasmic extracts, we found that proteins related to oxidative phosphorylation and integration of energy metabolism were overrepresented in CD4+ T cells from older compared to younger individuals. At the same time, gene expression analysis showed that pathways related to oxidative phosphorylation and the electron transport chain were downregulated and mitochondrial respiration was impaired in cells from older compared to younger participants. Interestingly, transmission electron microscopy showed no difference in the number of mitochondria in both naïve and memory CD4+ T cells from younger and older participants, but substantial differences of mitochondrial morphology. In particular, many of the mitochondria from naïve and memory CD4+ T cells from older participants were morphologically distorted and enclosed into autophagosome vesicles. Autophagic flux assays confirmed reduced autophagy efficiency, suggesting that in older persons defective autophagy may impair the recycling of irreversibly damaged mitochondria leading to buildup of OXPHOS proteins. We propose that interventions that normalize autophagy may slow down immunosenescence and prevent its deleterious consequences.
Results
Protein expression in human CD4+ T Cells
We examined the protein profiles from cytoplasmic extracts of human CD4+ T cells obtained from young (21-34 years-old) and older (68–83 years-old) male participants of the BLSA (Supplementary Table 1). Total CD4+ T cells were isolated from apheresis-derived peripheral blood mononuclear cells (PBMCs). Total CD4+ T lymphocytes, without distinction between naïve and memory subclass, were used for protein expression analysis because of the high number of cells needed to obtain enough cytoplastic extract to perform the proteomic study. The proportion of naïve CD4+ T cells (based on CD45RA+ expression) in the younger or older groups were comparable (Supplementary Figure 1), reflecting earlier published observations [28, 29]. Cytoplasmic extracts from CD4+ T cells were labeled with isobaric tags for relative and absolute quantification (iTRAQ) of proteins by liquid chromatography-mass spectrometry (LC-MS/MS). We compared 18 cytoplasmic extracts from older participants against four extracts from younger participants using four different iTRAQ 8plex kits. Equal amounts of cytoplasmic extract were used. One control sample from young participants was included in all four iTRAQ sets and used for protein quantitative normalization. To enhance comparability, technical controls (duplicate samples) were included within the same iTRAQ and also between iTRAQs (Supplementary Table 1). About 3,510 proteins were identified from each iTRAQ experiment. Data were analyzed within each iTRAQ 8plex and also in combination. Twenty-nine proteins (Table 1), ranked by significance (only proteins with p < 0.05 shown), were found to be up-regulated (ratio >1.3) in CD4+ T cells from older donors compared to younger donors after meeting additional analytical criteria (≥ 2 unique peptides, high confidence of the peptide sequences, and ion score of more than 30; see Materials and Methods). Interestingly, 23 of these were mitochondrial proteins (16 belonged to the electron transport chain; Figure 1A and Supplementary Table 2).
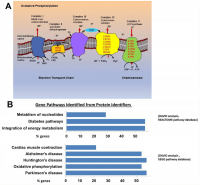
Figure 1. (A) Mitochondrial electron transport chain proteins that we found to be more highly expressed in the old are indicated in red by their location (Complex II, IV, and V) in the cartoon schema. (B) Gene pathways identified for proteins upregulated in the old using DAVID to convert protein identifiers to their genes (DAVID Bioinformatics Resources 6.7 (https://david.ncifcrf.gov). These genes fall into pathways related to oxidative phosphorylation, neurodegenerative disease, and integration of energy metabolism (Oxidative phosphorylation [16 from KEGG pathway database], Neurodegenerative disease [16 from KEGG], Cardiac muscle contraction [6 from KEGG], Integration of energy metabolism [16 from REACTOME pathway database], Diabetes [16 from REACTOME], Metabolism of nucleotides [8 from REACTOME]).
Using the Database for Annotation, Visualization and Integrated Discovery (DAVID), we found that proteins increased in CD4+ T cells from older participants mapped to several molecular pathways, namely oxidative phosphorylation (16 from KEGG pathway database), cardiac muscle contraction (6 from KEGG), integration of energy metabolism (16 from REACTOME pathway database), and metabolism of nucleotides (8 from REACTOME) (Figure 1B). These data demonstrate that mitochondrial proteins and particularly proteins that pertain to electron transport chain complexes accumulate with age in CD4+ T lymphocytes.
Gene expression in human CD4+ T Cells
To understand whether the higher accumulation of mitochondrial proteins in the cytoplasm of CD4+ T lymphocytes from older compared to younger participants was due to increased transcription/translation or rather to defective elimination, we studied gene expression in human CD4+ T cells by using the Parametric Analysis of Gene Enrichment (PAGE) analysis [29, 30] of normalized hybridization data from cRNA. Cells from 33 donors were analyzed divided into a young group (5 men and 3 women, ages 25–38 years-old) and an older group (19 men and 6 women, ages 70–82 years-old), (Supplementary Table 3). Expression pathways were transformed into Z-scores, and significant differences (p-value ≤ 0.05) between young (Y) and old (O) donors were identified (Figure 2, Supplementary Figure 2A, 2B).

Figure 2. Gene expression analysis showing top 51 up- and down-regulated biological pathways in CD4+ T cells between older (70 to 80 years-old) and younger (20 to 30 years-old) men. Mitochondria-related and oxidative phosphorylation-related pathways were the most down-regulated pathways in older compared to younger donors. Differences of Z-scores between younger and older participants are shown on the X-axis. Each row denotes a different pathway (p ≤ 0.05 and FDR ≤ 0.3). N = 5 young, 19 old donors.
To be consistent with proteomic data, we first compared gene expression only between young and old men. Contrary to our proteomic analysis, PAGE identified “mitochondria-related” and “oxidative phosphorylation-related” pathways were amongst the most down-regulated biological pathways in older compared to younger individuals (Figure 2). That is, for many proteins found to be overrepresented in older persons CD4+ T lymphocytes, gene transcripts were instead underrepresented in CD4+ T lymphocytes from older persons. The same trends were noted when the analysis was restricted to women or included both sexes (Supplementary Figure 2A, 2B). Thus, proteomic and gene expression results tended to be inversely correlated.
Differences of mitochondrial respiration between young and old CD4+ T Cells
To understand whether overrepresentation of mitochondrial proteins in CD4+ T lymphocytes from older compared to younger individuals comes from functional or dysfunctional mitochondria, we measured mitochondrial respiration in CD4+ T cells of 7 young (ages 22–35 years-old) and 7 old (ages 80–93 years-old) male participants (Supplementary Table 4A) using high-resolution respirometry OROBOROS Oxygraph-2k (O2k). We challenged CD4+ T lymphocytes from younger and older participants with several mitochondrial inhibitors in sequence to estimate mitochondrial reserve capacity, nonmitochondrial oxygen capacity, ATP-linked respiration and proton leak and we used these values to derive the Bioenergetic Health Index (BHI), a summary index of mitochondrial function in cells [31]. The BHI is derived from calculating a ratio of positive aspects of mitochondrial bioenergetic function (i.e. reserve capacity and ATP-linked respiration) to potentially deleterious aspects of mitochondrial bioenergetic function (i.e. non-mitochondrial oxygen consumption and proton leak) [31].
BHI was higher in cells from younger compared to older participants (Supplementary Figure 3, representative mitochondrial respiration measurements), except for one outlier whose flow cytometric profile was different from the other 13 samples and the percentage of memory cells (central and effector) much higher than that percentage in the other 6 older donors (59% compared to a range 26–48%) (Figure 3A and Supplementary Table 4B). After accounting for differences due to different percent of memory cells between groups, the BHI differences between cells from younger and older subjects was statistically significant (p = 0.036) (Figure 3B). Of note, the majority (85%) of CD4+ T cells from younger participants had higher BHI compared to cells from older donors. Conversely, nonmitochondrial respiration, typically attributed to the action of cyclooxygenases, lipoxygenases and NADPH oxidases that are considered negative indicators of bioenergetic health, was significantly higher (p = 0.049) in CD4+ T lymphocytes from older compared to younger participants [31] (Figure 3C). We also found that the reserve mitochondrial capacity was significantly higher (p = 0.045) in cells from younger compared to older participants (Figure 3D). Cumulatively, these observations demonstrate that despite increased abundance of mitochondrial proteins, mitochondrial function of CD4+ lymphocytes is lower in older compared to younger individuals.
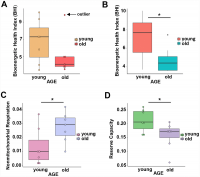
Figure 3. (A) Calculated bioenergetic health index (BHI) from young and old CD4+ T cells. The BHI is derived from calculating a ratio of positive aspects of mitochondrial bioenergetic function (i.e. reserve capacity and ATP-linked respiration) to potentially deleterious aspects of mitochondrial bioenergetic function (i.e. non-mitochondrial oxygen consumption and proton leak). Cellular mitochondrial function was determined using high-resolution respirometry with oligomycin, FCCP, rotenone, and antimycin A. For BHI, one outlier of 14 participants was noted and a comparison of the old and young BHI with this outlier was not significantly different (p=0.19). (B) In further analysis, FACS showed the outlier subject had a higher percentage of total memory CD4+ T cells (59% compared to a range 26-48%) than other participants. Adjusting for the average percentage of memory CD4+ T cells, the calculated the BHI was significantly higher for younger compared to older participants (*p = 0.036). © Nonmitochondrial respiration was found to be significantly higher in CD4+ T cells from older compared to younger individuals (*p = 0.049). (D) Reserve capacity was significantly higher in cells from young compared to older participants (*p = 0.045). (A–D) P-values were calculated by Welch’s t-test, a variation of the Student’s t-test that does not make the assumption of equal variance in the two compared samples [55]. Error bars reflect the standard error of the mean (±SEM). N = 7 young, 7 old donors.
.../...
F O R T H E R E S T O F T H E S T U D Y, P L E A S E V I S I T T H E S O U R C E .
.
Edited by Engadin, 19 November 2019 - 04:53 PM.
















































