.
S O U R C E : Max Planck Institute for Biology of Ageing
P A Y W A L L E D P R I M A L S O U R C E : nature (NFYB-1 regulates mitochondrial function and longevity via lysosomal prosaposin)
As people get older, they often feel less energetic, mobile or active. This may be due in part to a decline in mitochondria, the tiny powerhouses inside of our cells, which provide energy and regulate metabolism. In fact, mitochondria decline with age not only in humans, but in many species. Why they do so is not well understood. Scientists at the Max Planck Institute for Biology of Ageing in Cologne set out to understand how mitochondrial function is diminished with age and to find factors that prevent this process. They found that communication between mitochondria and other parts of the cell plays a key role.
Microscopy image of a C. elegans worm (red: nuclei, in which NFYB-1 is present, green: lysosomes). © Raymond Laboy
For their studies, the scientists used the simple roundworm, Caenorhabditis elegans, an important model system for ageing research. Over half the genes of this animal are similar to those found in humans, and their mitochondria also decline with age. From their research, the scientists found a nuclear protein called NFYB-1 that switches on and off genes affecting mitochondrial activity, and which itself goes down during ageing. In mutant worms lacking this protein, mitochondria don’t work as well and worms don’t live as long.
Unexpectedly, the scientists discovered that NFYB-1 steers the activity of mitochondria through another part of the cell called the lysosome, a place where basic molecules are broken down and recycled as nutrients. “We think the lysosome talks with the mitochondria through special fats called cardiolipins and ceramides, which are essential to mitochondrial activity,” says Max Planck Director, Adam Antebi, whose laboratory spearheaded the study. Remarkably, simply feeding the NFYB-1 mutant worms cardiolipin restored mitochondrial function and worm health in these strains.
Because cardiolipins and ceramides are also essential for human mitochondria, this may mean human health and ageing can be improved by understanding on how such molecules facilitate communication between different parts of the cell. This work has been recently published in Nature Metabolism.
Mitochondrial activity is critical for cellular vitality and organismal longevity, yet underlying regulatory mechanisms in metazoans remain elusive. To identify mitochondrial regulators, they performed an RNAi screen leveraging the remarkable mitochondrial changes in C. elegans upon recovery from adult reproductive diapause. They discovered NFYB-1, a subunit of the NF-Y transcriptional complex, as a crucial regulator of mitochondrial function. Loss of NFYB-1 leads to reduced mitochondrial gene expression, mitochondrial fragmentation, and abolition of longevity triggered by mitochondrial impairment. Moreover, NFYB-1 deletion disrupts mitochondrial UPRmt factors and mitochondrial-to-cytosolic stress response (MCSR). Multi-omics analysis indicates that NFYB-1 serves as a potent repressor of several ER genes and the ER stress response, as well as lysosomal prosaposin. Downstream of NFYB-1, limiting prosaposin expression alters ceramide and cardiolipin pools, restores mitochondrial fusion, gene expression and longevity. Thus, the NFYB-1/PSAP axis coordinates lysosomal to mitochondrial communication to prolong life.
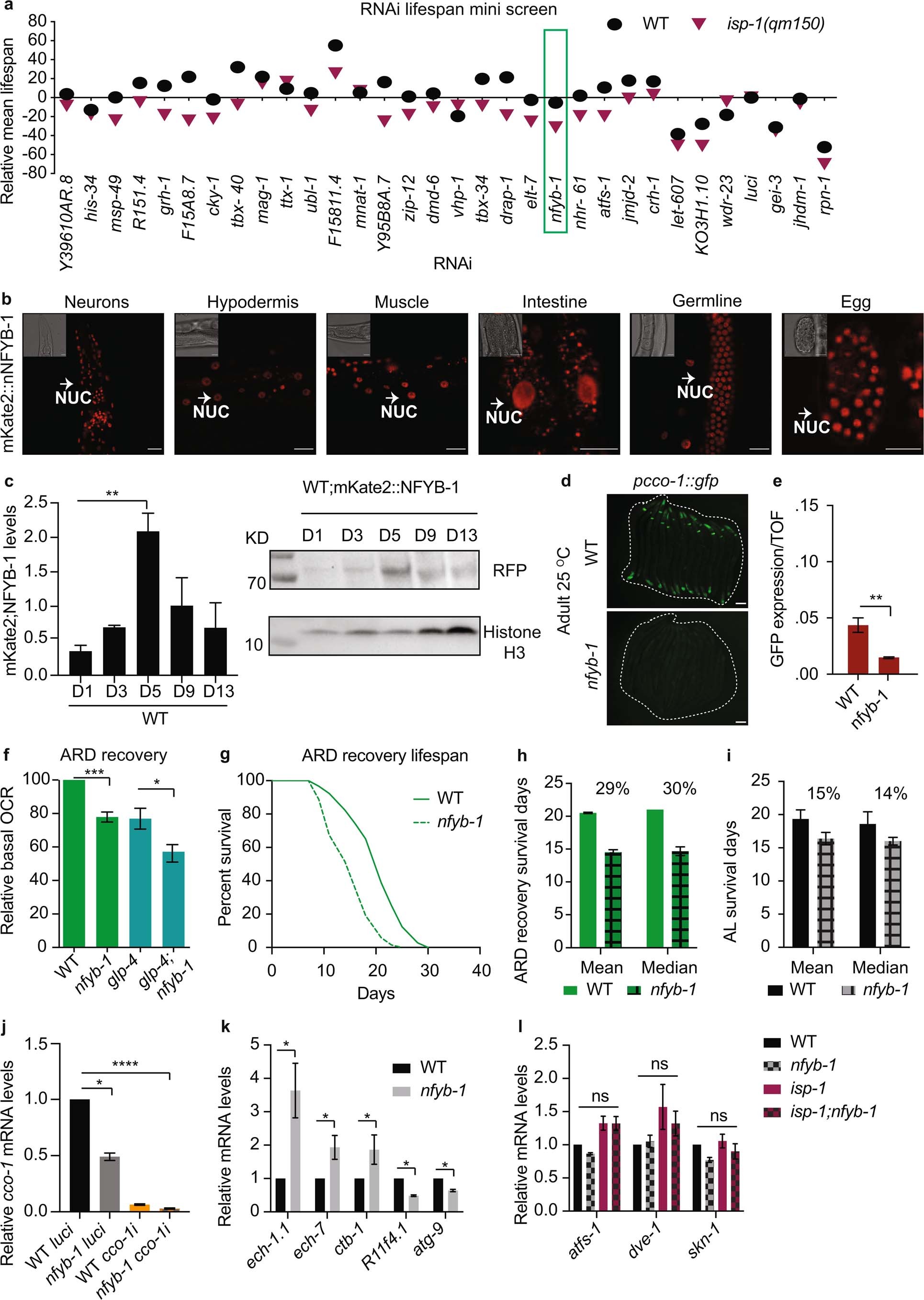
Extended Data Fig. 1: NFYB-1 regulates mitochondrial function and longevity (Related to Fig. 2).
a, Mini lifespan screen of WT and isp-1(qm150) upon RNAi knockdown of candidate genes from egg on in comparison to luci RNAi control bacteria (N = 1, n = 120 per condition). b, Representative image of mKate2::NFYB-1 nuclear (nuc) localization in various tissues of AL day 1 adults, scale bar = 10 μm. c, Representative immunoblot of mKate2::NFYB-1 levels over age in AL WT (day 1, 3, 5, 9 and 13 as mentioned, n = 200 worms per repeat per condition), with quantitation and histone loading control. d-e, nfyb-1(cu13) mutations leads to a decrease in pcco-1::gfp expression in AL day 1 at 25 °C. d, Representative images of ca. n = 10–20 worms, scale bar = 75 μm, e, pcco-1::gfp expression levels normalised to (Time of Flight) TOF quantitated using biosorter (n ≥ 400). f, nfyb-1(cu13) mutation results in a decrease in relative basal oxygen consumption rate (OCR) compared to WT, and in germline-less glp-4(bn2);nfyb-1(cu13) compared to glp-4(bn2) animals at 25 °C, upon ARD recovery (n = 900 worms). g, Kaplan–Meier survival curves of WT, nfyb-1(cu13) upon ARD recovery shows that nfyb-1 loss leads to decrease in survival (n = 120 per repeat, per condition). h, i, Percentage decrease in mean and median survival of nfyb-1(cu13) in comparison to WT in AL and upon ARD recovery. j, cco-1 mRNA levels are reduced in nfyb-1(cu13). k, mRNA levels of nfyb-1 mitochondrial regulated genes, l, loss of nfyb-1 does not affect mRNA levels of atfs-1, dve-1 and skn-1, j-l quantitation by qPCR at AL day 1 (n ≥ 1000). b-l, All data represent N = 3 independent biological replicates, n = total worms per condition from three replicates unless stated otherwise. Error bar shows mean± s.e.m., statistics determined by c, j, l one-way ANOVA and e, f, k two-sided t-test, ns: not significant, *P < 0.5, **P < 0.01, ***P < 0.001. a, g-i, Two-sided Mantel–Cox log-rank test, refer to Supplementary Table 2 for statistics. d-f, ARD day 10 worm recovered for 1day. Source Data
.
Edited by Engadin, 21 May 2020 - 10:41 PM.















































