.
O P E N A C C E S S S O U R C E : AgING
Abstract
Bats are the longest-lived mammals given their body size with majority of species exhibiting exceptional longevity. However, there are some short-lived species that do not exhibit extended lifespans. Here we conducted a comparative genomic and transcriptomic study on long-lived Myotis myotis (maximum lifespan = 37.1 years) and short-lived Molossus molossus (maximum lifespan = 5.6 years) to ascertain the genetic difference underlying their divergent longevities. Genome-wide selection tests on 12,467 single-copy genes between M. myotis and M. molossus revealed only three genes (CCDC175, FATE1 and MLKL) that exhibited significant positive selection. Although 97.96% of 12,467 genes underwent purifying selection, we observed a significant heterogeneity in their expression patterns. Using a linear mixed model, we obtained expression of 2,086 genes that may truly represent the genetic difference between M. myotis and M. molossus. Expression analysis indicated that long-lived M. myotis exhibited a transcriptomic profile of enhanced DNA repair and autophagy pathways, compared to M. molossus. Further investigation of the longevity-associated genes suggested that long-lived M. myotis have naturally evolved a diminished anti-longevity transcriptomic profile. Together with observations from other long-lived species, our results suggest that heightened DNA repair and autophagy activity may represent a universal mechanism to achieve longevity in long-lived mammals.
Introduction
Natural selection has shaped a large variation of lifespan across mammals, with maximum lifespan ranging from a few months (e.g. short-lived shrews) to 211 years (e.g. bowhead whale) [1]. Although the bowhead whale is exceptionally long-lived, its lifespan is arguably not as extreme as that of a 30 years old naked mole-rat given their body sizes, as maximum lifespan (MLS) exhibits a positive correlation with body size within mammals [2, 3]. Thus, lifespan comparison across mammals requires body size correction. To resolve this, the longevity quotient (LQ) was introduced, which is defined as the ratio of observed lifespan to predicted lifespan for a non-flying mammal of the same body size [2, 3]. Using this approach bats are the longevity extremists, with some species living up to ten times longer than expected given their body size [2]. The Brandt’s bat (Myotis brandtii) holds the record for longevity [4], with a maximum lifespan of >40 years, living 8~10 times longer than expected given body size (~7 grams) [2, 4, 5]. This renders bats as one of the most ideal taxa to explore the molecular basis of extraordinary longevity in mammals.
Although the majority of bat species are long-lived, especially within the Myotis genus, there are a few short-lived exceptions, such as the velvety free-tailed bat (Molossus molossus) and the evening bat (Nycticeius humeralis), living as long as would be expected given their body size [5, 6]. A recent study has suggested that the ancestral bat lived up to 2.6 times longer than expected given body size, indicating that the extreme longevity observed in the longest-lived bat genera may have evolved multiple times [7]. This also suggests that short-lived bat species may have lost their longevity adaptations. Therefore, this wide range of lifespans observed in bats enables us to utilize comparative evolutionary approaches to search for genetic differences within closely-related long- and short-lived bat species. In contrast to comparative studies on phylogenetically-distant species (e.g. bats versus mice), this comparison could minimize the ‘noise’ resulting from heterogenous physiology, and may reveal key anti-aging molecular adaptations that have evolved in long-lived bats, or were lost in their short-lived counterparts.
Genome-wide comparative analyses have been carried out on a few long- and short-lived species in primates [8], rodents [9, 10], whales [11] and bats [12]. These studies revealed a few genes that showed convergent amino acid mutations, or exhibited positive selection, or were differentially expressed in long-lived species compared with their short-lived counterparts. Although there is little commonality across the gene candidates that are associated with longevity revealed by these studies, they are mainly enriched in DNA repair and maintenance, autophagy, homeostasis, and nutrient sensing pathways [13]. In bats, our previous longitudinal studies showed that long-lived M. myotis bats maintained their transcriptomic profiles [14] and telomere length [5], and did not exhibit an increased level of mitochondrial damage with advancing age [15], all of which likely contribute to their extraordinary longevity. However, a parallel comparison between long- and short-lived bats is lacking.
In this study we performed a comparative genomic and transcriptomic analysis between long-lived Myotis myotis (MLS = 37.1 years; LQ = 5.71) and short-lived Molossus molossus (MLS = 5.6 years; LQ = 0.99) [6] to ascertain the molecular signatures associated with longevity in bats. Based on the genome-wide alignments of single-copy orthologous genes between these two species, we detected and further investigated the genes that were fast-evolving and showed significant positive selection. We also deep sequenced blood transcriptomes from eight adult individuals for each species, and explored the genes and pathways that were differentially expressed. To ascertain if long-lived bats have evolved a transcriptomic signature of longevity, we further investigated the expression of ‘pro’- and ‘anti’-longevity genes in the blood transcriptomes of M. myotis and M. molossus. Although the majority of genes underwent purifying selection, we observed a significant transcriptional alteration between these two species. Among 2,086 genes that exhibited large interspecific expression variation, the genes that showed higher expression in long-lived M. myotis were mainly enriched in DNA repair and autophagy. Further pathway analysis suggested that six biological processes, including autophagy, were differentially expressed between M. myotis and M. molossus. We also show that M. myotis had significantly lower expression levels of anti-longevity genes, suggestive of a transcriptomic signature of longevity naturally evolved in long-lived bats. Together with the previous findings in other long-lived mammals, our study implies that enhanced DNA repair and autophagy activity may represent a universal mechanism to achieve longevity in long-lived mammals.
Results
The majority of genes undergo purifying selection
To evaluate the natural selection acting on protein-coding genes between M. myotis and M. molossus, we calculated the ratio of Ka and Ks substitution rates for each pair of orthologous genes. We observed that most of the genes (97.96%) were under purifying selection (Ka/Ks ratio < 1, FDR < 0.05), with the median of Ka/Ks ratios equal to 0.103 (Figure 1A). In total, 48 genes had the ratios of Ka/Ks > 1, only three of which (FATE1, MLKL and CCDC175) exhibited significant positive selection (Fisher’s exact test, FDR < 0.05, Figure 1B). To determine if positive selection on these genes was the consequence of species-specific selective pressures, we conducted pairwise analyses of orthologous genes across 6 bat species (See Methods). We found that each comparison resulted in a unique set of positively selected genes (PSGs) (Supplementary Table 1). Interestingly, positive selection on FATE1 and CCDC175 was also observed between M. molossus, M. myotis and other bat species (Supplementary Table 1). Although MLKL was solely seen under significant positive selection between M. myotis and M. molossus, this gene also showed high Ka/Ks ratios between all possible comparisons of bat species (Figure 1C and 1D).
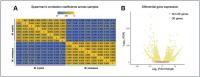
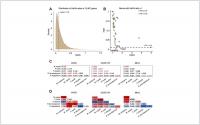
Figure 1. Analysis of Ka/Ks substitution rates of 12,467 single-copy genes between M. myotis and M. molossus. (A) Distribution of Ka/Ks ratios of 12,467 single-copy genes. To better visualize the distribution, six genes with Ka/Ks > 1.5 were not included in this plot. (B) Genes with Ka/Ks > 1. Three genes highlighted in red show significant positive selection (Ka/Ks > 1; FDR < 0.05 Fisher’s exact test). © Significance (FDR) of Ka/Ks ratios of FATE1, CCDC175 and MLKL between 6 bat species through pairwise comparisons. The red values indicate significant positive selection while the blue values indicate significant purifying selection. The black values indicate no selection. (D) Ka/Ks ratios of FATE1, CCDC175 and MLKL between 6 bat species through pairwise comparisons. The red values indicate Ka/Ks ratios > 1 while the blue values indicate Ka/Ks ratios < 1.
Transcriptomic profiles exhibit a substantial difference in long-lived and short-lived bats
To ascertain the differences on the transcriptional level, we further sequenced and compared their blood transcriptomes (n = 16) using Illumina RNA-Seq. Out of 12,467 orthologs, we excluded 1,832 genes that were not expressed in either M. molossus or M. myotis and retained 10,635 genes for downstream analyses. The correlation analysis revealed a considerable difference in global transcriptomic profiles between M. molossus and M. myotis (Figure 2A, See Methods). The samples showed statistically higher intraspecific Spearman’s correlation coefficients (ρ = 0.957) in contrast to the interspecific counterparts (ρ = 0.766) (P = 2.2×10-16, Wilcoxon signed-rank test). In addition, 4,846 (45.57%) genes were differentially expressed (FDR < 0.05). 2,162 (20.33%) genes showed up-regulation in M. myotis compared to M. molossus while 2,684 (25.24%) genes were down-regulated (Figure 2B). Interestingly, there was no significant difference in the distribution of Ka/Ks ratios between differentially and non-differentially expressed genes (P = 0.162, Kolmogorov-Smirnov test). Next, we tested if the Ka/Ks ratios of 10,635 orthologs had an association with their differential expression patterns in M. molossus and M. myotis. We found that there was no significant association between the genes differentially expressed and their Ka/Ks ratios (P = 0.352, χ2 test, See Methods).
Figure 2. Comparisons of M. myotis and M. molossus blood transcriptomes. (A) Spearman’s correlation coefficients between M. myotis and M. molossus blood transcriptomes based on expression levels of 10,635 single-copy genes. We excluded 1,832 genes that were neither expressed in M. myotis nor M. molossus. (B) Differential gene expression analysis between M. myotis and M. molossus blood transcriptomes. Genes with FDR < 0.05 were considered differentially expressed genes (DEGs).
DNA repair and autophagy are enriched by the genes that have genetically higher expression in long-lived bats
For each of 10,635 genes, we calculated the proportion of interspecific expression variation using a linear mixed model (See Methods). On average, 24.9% of gene expression variance was explained by ‘species’ while ‘sex’ only explained a small proportion of variation (Figure 3A). To investigate the gene expression that genetically differed in M. myotis and M. molossus, we focused on 2,086 genes with at least 80% of their expression variance resulted from ‘species’ which represented interspecific variation (See Methods). As was expected, 2,083 of these genes (99.86%) were also detected as differentially expressed genes (DEGs) (Figure 3B).
.../...
.