.
Alexander Fedintsev worked at the Research Institute of Antimicrobial Chemotherapy (Smolensk, Russia). Now he is an engineer in machine learning systems. Biogerontologist scientist specialized in biostatistics and data analysis.
F U L L T E X T S O U R C E : Medium (Note: Original in russian. Google translated).
Horvat’s epigenetic clock is probably the most famous “achievement” of bioherontology (in the second part of the cycle, I analyze in detail the reasons why the word achievement is in quotation marks). Nevertheless, they accurately predict the passport age and it is this property that has generated many myths around itself. At times, completely miraculous properties that they do not possess are attributed to the epicases. Let's try to make out some of the myths around the Horvat watch and understand what processes are behind age-related epigenetic changes.
Let's start with the popular misconception: “Horvat watches show almost the same “time” in neurons, and in lymphocytes, and in dozens of other tissues.”
The epichase error for buccal (on the inside of the cheek) epithelium is less than a year, while for heart tissue it is 12 years, for the mammary gland 13 years, and for skeletal muscles - 18 years! Well, where is “almost the same time” here? That is, a difference of 20 times is not significant !? This is about how to say that you are 5 minutes late, and arrive in an hour and a half.
The second misconception: the hypothalamus controls the “epigenetic program of aging”
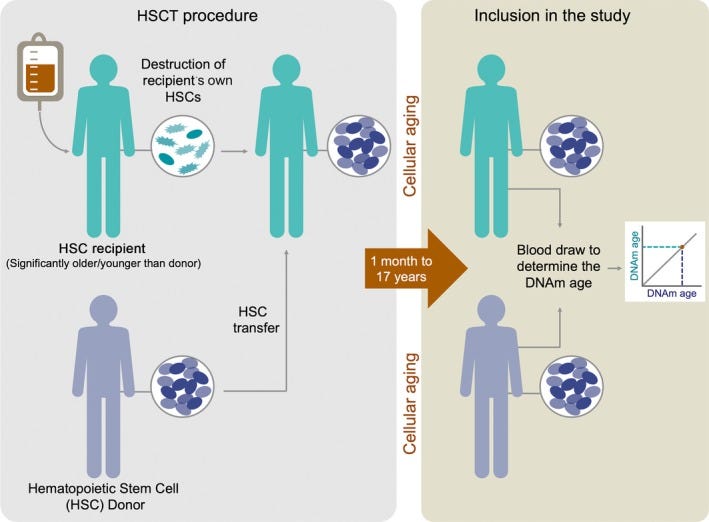
Moreover, the epigenetic age is most likely an intrinsic property of the cells and does not depend on the environment: in the study, recipients were killed by their stem blood cells, transplanted by donor cells and looked at the correlation of the age of blood cells with the ages of the donor and recipient:
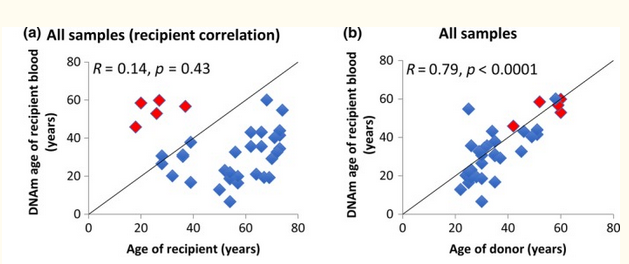
And it was extremely surprising that the epigenetic age of the transplanted cells even after years corresponded to the age of the donor, and not the recipient! Obviously, neither the hypothalamus, nor anything else externally affects epigenetics!.
One can see the absence of correlation of the age of the cells with the age of the recipient and a rather strong correlation with the age of the donor.
At the same time, we know that young medium allows stem cells to live much longer than the maximum species life span, and the old medium, accordingly, ages young cells. Therefore, a reasonable question arises: does the epigenetic age reflect the aging of the body? I examined this issue in more detail in my last article.
The third misconception: epigenetic changes during aging are not random, but programmed
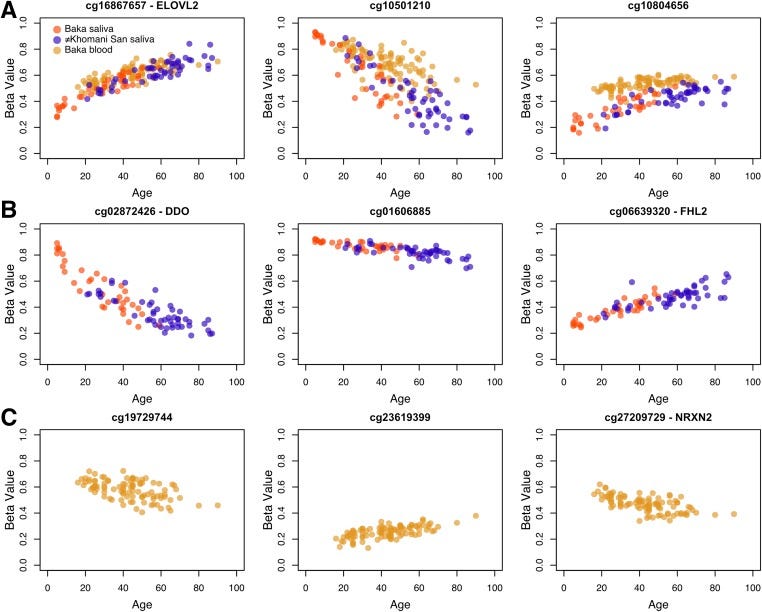
We will dwell on this myth in more detail. To begin with, the methylation level is not a deterministic quantity . It is itself stochastic! Take a look at the chart from a great article :
Graph of methylation changes for some sites with age.
We see how methylation of various sites (those that are most significantly associated with age) changes with age. The vertical axis of each chart is the Beta Value. Roughly speaking, this is the proportion of cells of the same tissue with a methyl tag in a certain position. And pay attention, this share is not equal to 100% or 0%! That is, even at the level of cells of one tissue there is a scatter: in some cells a certain site is noticed, while in others it is not. And scatter always means the presence of a stochastic component. Plus there is a spread between people. That is, the presence of stochastics in epigenetic changes during aging is a fact, but the presence of any programmed changes still needs to be proved. At the same time, the principle of “Occam’s razor” operates in science - that is, preference is given to simpler hypotheses, which describe the observed data and do not require the involvement of new entities. Obviously, a certain program is a new entity, so the question arises: is it possible that changes in methylation are purely stochastic in nature?.
Pay attention to one very interesting detail: sites where methylation was initially high tend to demethylate and vice versa. This fits very well into the following logic: if a methyl mark is present, then it can only fly off (if we are talking about a random process). And vice versa: if there is no methyl tag, then it, accordingly, may appear with some probability. But what random processes can lead to methylation and demethylation? And there are at least two such processes:
DNA repair. So, it has long been known that excision repair of bases (works in the cell in case of single-stranded DNA damage) actively removes methyl marks . The Gadd45 protein, which plays an important role in DNA repair, is also involved in demethylation . Interestingly, demethylation at the zygote formation stage is also associated with DNA repair! But not only demethylation is associated with DNA repair. For example, ultraviolet DNA damage leads to increased methylation of the CHK1 gene. In addition, it was shown that it is the repair of double-stranded DNA breaks that leads to an increase in methylation (not only of cytosines, but also of histones ).An elegant experiment showed that homologous recombination during DNA repair changes methylation and modifies histones, which leads to different levels of expression of the fluorescent GFP protein in genetically identical cells. Well, just like in our case with the cells of one tissue! A recent experiment by Sinclair and co-authors perfectly summed up all previous studies: they showed that the induction of non-mutagenic double-stranded breaks leads to methylation changes and, at the same time, to the accelerated aging phenotype. This alone is enough, but there is something else.
Krebs cycle metabolites. In our energy stations - mitochondria - this wonderful cycle of tricarboxylic acids is spinning , thanks to which, ultimately, the cell’s energy currency, ATP, is synthesized. So it turns out that one of the metabolites of the cell - alpha-ketoglutarate - is a substrate of demethylating enzymes such as JmjC and TET. And succinate and fumarate, on the contrary, lead to hypermethylation . Thus, disruptions in the functioning of the citrate cycle can lead to an imbalance of its metabolites in the cytoplasm and corresponding changes in methylation. And about the Krebs cycle, you still see the video, it is useful to know.
Thus, absolutely all types of epigenetic changes (both methylation and demethylation) during aging can be explained by stochastic processes (DNA damage, impaired mitochondria). This study is also in favor of this study (thanks to Denis Odinokov for the
link).
Here, the authors observed 375 twins for a long time (18 years). Researchers not only once measured methylation levels, but did it many times (at least 5 times). The fact that the study was conducted on the twins tells us that if there was an aging program, it would work the same way for the twins - that is, their methylation would change in a very similar way. But this was just not discovered, both in this study and in others!
Some may argue: what if most methylation sites change randomly and a small subset is programmed? The most important candidate for such a subset is the 353 methylation sites on which the very Croat watch is based. But here, too, supporters of the program will be disappointed: the study showed that the heritability of the difference between passport age and epiclock readings is rather weak and amounts to only 0.43. Hannum watches have approximately the same heritability, so the program is most likely not there.
Moreover, there are longitudinal studies , where they looked at the correlation of twin epi-ages in time:
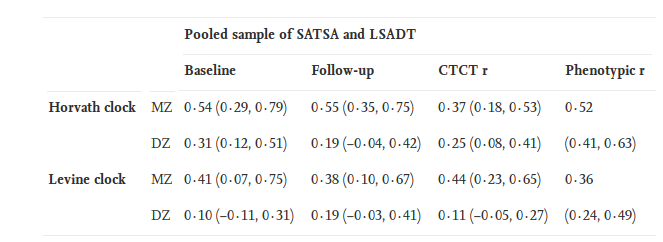
What do we see in the table? We see that the correlation between the epi-ages of identical twins is rather low — a maximum of 0.55. Even weaker are these correlations in Levin’s watches. And this despite the fact that identical twins have exactly the same genome and extremely similar epigenes at the start! If the methylation of the sites of Horvat watches changed according to some program, then I would expect to see a correlation between identical twins in the region of 0.85-0.9, but not at all 0.55 (the IQ correlation coefficient between twins is 0.86, and the growth correlation coefficient is 0.79–0.93, in average 0.88). And here is a study, which shows that although identical twins are identical epigenetically in infancy, in adulthood they are already very significantly different from each other, which indicates that epigenetic changes are the result of the combined influence of random processes and the environment.
.../...
.
Edited by Engadin, 18 December 2019 - 06:04 PM.